Blog
P4 Allele of MYOZ3
Introduction
This blog post explores the P4 allele (S42L) of the equine MYOZ3 gene, which encodes myozenin 3. Portions of this blog post serve as additional sources of information to supplement the MYOZ3 Gene Page.
We present data to support the hypothesis that the P4 allele of MYOZ3 (S42L) is damaging.
The substitution of leucine (hydrophobic) for serine (polar uncharged) in the P4 allele of MYOZ3 is a nonconservative substitution of a chemically dissimilar amino acid.
Evolutionary conservation provides convincing evidence that the P4 allele of MYOZ3 is damaging. We use public data to show that the reference allele is completely conserved across 448 species of mammals and birds representing 155 unique sequences of the 41-amino acid query sequence centered on the S42L allele, covering over 310 million years of evolutionary history.
P4 allele of equine MYOZ3
Evolutionary conservation provides evidence on whether the P4 allele of MYOZ3 is damaging. In this approach, predicted MYOZ3 protein sequences are compared among a number of different species. This method, applied to mammals and birds, is shown below.
A 41 amino acid segment of the protein sequence of equine MYOZ3 (XP_005599348.1), centered on the position of the variant, is shown below. This sequence is compared to a segment of the protein sequence of the equine P4 allele (S24L). The affected amino acid is highlighted in red.
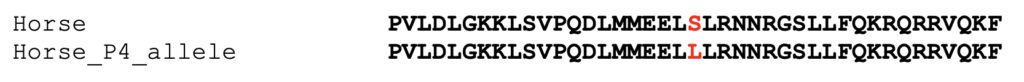
The 41 amino acid portion of the equine MYOZ3 gene centered on the variant, shown above, was used to conduct a blastp search of the protein sequence database at NCBI, retrieving only sequences from mammals and birds. Identical sequences were clustered. Sequences represented by only a single species were used as query sequence for additional blastp searches.
Evolutionary Conservation: Mammals
Evolutionary conservation provides evidence on whether the P4 allele of MYOZ3 is damaging. In this approach, predicted MYOZ3 protein sequences are compared among a number of different species. This method, applied to mammals, is shown below. After clustering of identical sequences, the sequences from 160 mammals represent 37 unique sequences. These were aligned using CLUSTAL as shown in Figure 1.

Figure 1. Alignment of partial MYOZ3 protein sequences from mammals. The horse sequence was used as a blastp query sequence to retrieve MYOZ3 protein sequences from mammals. Sequences that were identical were clustered. Numbers in parentheses indicate the number of species in a cluster. CLUSTAL output summarizes whether a particular position is a single and fully conserved residue (*), has a conservative substitution with strongly similar properties (:), a somewhat conservative substitution (.), or is not conserved ( ). The sequence of the horse P4 allele is shown for comparison, but was not included in the CLUSTAL analysis. The position of S42L is highlighted in red. See the Technical Appendix for details.
The alignment shown in Figure 1 demonstrates that the S42 allele is invariant throughout the mammalian lineage; no other amino acid has gone to fixation at this position in any species. Conservation of the S232 allele is seen throughout mammals, including marsupials (wombat, koala, and some marsupials included in the mammalian clusters) and monotremes (platypus, echidna).
In the alignment shown in Figure 1, only 25 of the 41 positions are invariant throughout the mammalian lineage; the S42 allele is invariant. This shows that the equine MYOZ3-S42L allele is not tolerated over evolutionary time in mammals.
Evolutionary Conservation: Birds
Partial MYOZ3 protein sequences were recovered from 288 species of birds as described above. The sequences from 288 species represent 118 unique sequences. These were aligned using CLUSTAL as shown in Figure 2.

Figure 2. Alignment of partial MYOZ3 protein sequences from birds. The horse sequence was used as a blastp query sequence to retrieve MYOZ3 protein sequences from birds; bird sequences were used as queries to retrieve additional sequences. Sequences that were identical were clustered. Numbers in parentheses indicate the number of species in a cluster. CLUSTAL output summarizes whether a particular position is a single and fully conserved residue (*), has a conservative substitution with strongly similar properties (:), a somewhat conservative substitution (.), or is not conserved ( ). The sequence of the horse P4 allele is shown for comparison, but was not included in the CLUSTAL analysis. The position of S42L is highlighted in red. See the Technical Appendix for details.
In the alignment shown in Figure 3, only 19 of the 41 positions are invariant throughout the avian lineage; the S42 allele is invariant. This shows that the MYOZ3-S42L allele is not tolerated over evolutionary time.
Summary
The evidence presented here strongly supports the hypothesis that the MYOZ3-S42L allele found in horse is not tolerated over evolutionary time. The only occurrence of the S42L allele is as a minor allele in horse, a species not subject to natural selection. Sequence conservation at this position is absolute across 448 species of mammals and birds representing 155 unique sequences of the 41-amino acid query sequence centered on the S42L allele.
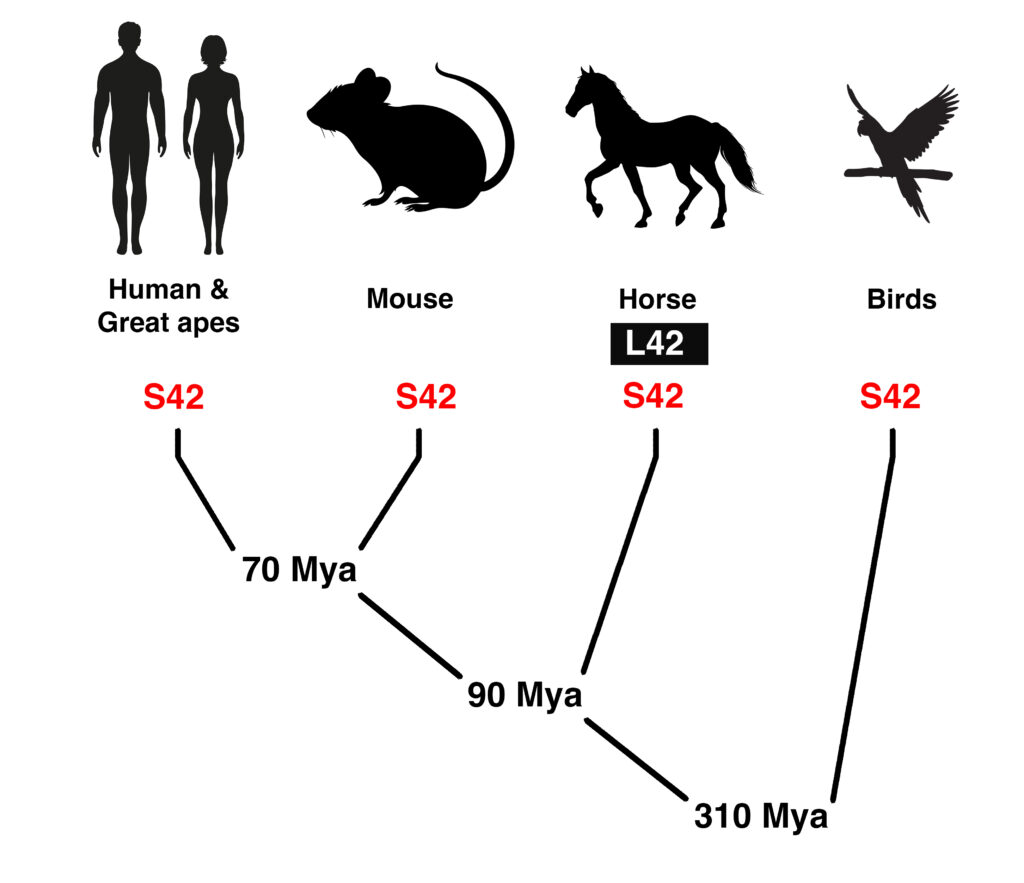
Figure 3. The amino acid at the position of the equine P4 allele of MYOZ3 (S42L) is a serine conserved throughout evolution. Sequence conservation at this position is absolute across 448 species of mammals and birds representing 155 unique sequences of the 41-amino acid query sequence centered on the S42L allele. Estimated time of divergence is shown as millions of years ago (Mya).
Technical Appendix
The purpose of this technical appendix is to permit researchers to reproduce and extend these results independently.
The equine MYOZ3-S42L allele, described by the coordinates and base substitution in EquCab 3.0, is chr14:26,710,261 G/A. The best protein model of equine MYOZ3 is XP_005599348.1. Click the link to the UCSC Genome Browser to view the horse genome sequence centered on the position of the S42L allele.
Retrieving protein sequences. Protein sequences like those shown in the alignments (Figures 1 and 2) can be retrieved from NCBI using the blastp tool and a query sequence.
The partial equine MYOZ3 query sequence used to retrieve sequences is:
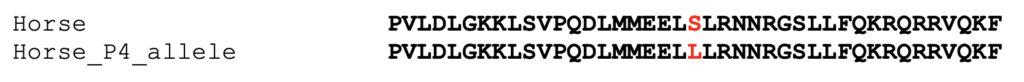
This sequence can be used as a query to identify related sequences using blastp as shown in Figure 4.
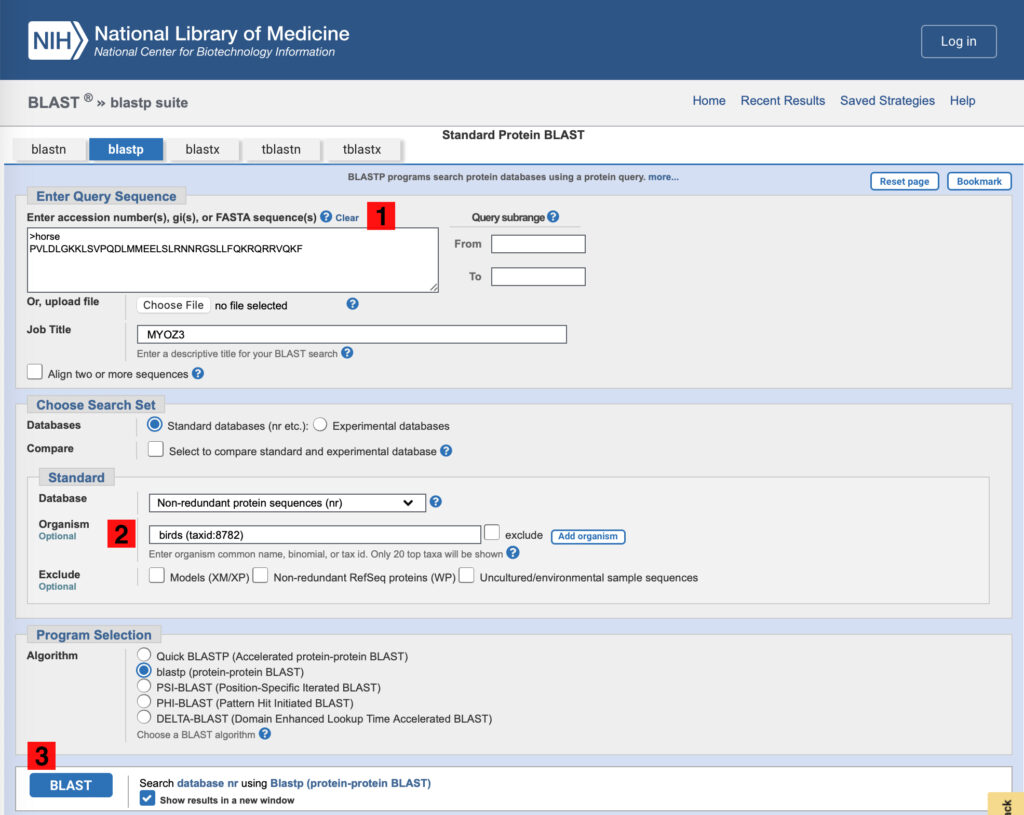
Figure 4. NCBI blastp search page. To use the equine MYOZ3 query sequence to identify MYOZ3 sequences from birds, 1) copy the equine sequence to the query sequence box, 2) set the optional Organism to “birds.” Note that this recovers a taxonomic ID as shown, and 3) Click BLAST.
Results for one species are shown below.
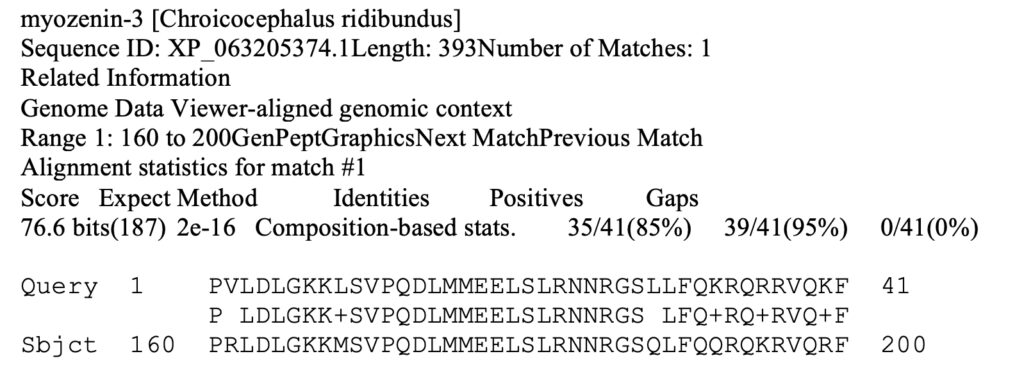

The match is to the Black-headed gull (Chroicocephalus ridibundus). The Query line shows the part of the equine sequence that matched the bird sequence. The Subject line shows the bird sequence. At each position, if the amino acid is identical in the two sequences, the amino acid is entered in the middle line. Conservative substitutions are marked with a “+” sign; nonconservative substitutions are left blank.
The recovered sequence can be put into FASTA format:

This can then be used as a query sequence for an additional blastp search.
Aligning protein sequences
Multiple protein sequences were aligned using CLUSTAL.
Evolutionary relationships
Information on evolutionary relationships among species is presented graphically as the Tree of Life.
Download data
The data used for alignments in Figures 1 and 2 are available as a downloadable spreadsheet.
For each sequence used in the analysis, the spreadsheet contains the figure in which the sequence appears. Each individual species in a cluster is identified by species name and common name. The sequence ID for the MYOZ3 protein sequence in that species is also shown.
Share this post
From the blog
The latest industry news, interviews, technologies, and resources.